| |
<Prev Next> |
In this example we will show some of the considerations required when fitting multiple states simultaneously, particularly with regards to the putting the line lists together for fitting. We start where the single state fit in Fitting example - the A-X transition in C3 finished by adding another state. The experimental spectrum does not seem to show the presence of another Q branch, suggesting a Σ-Σ transition should be added, rather than another Π-Σ transition like the first. If we assume the lower state is the same for both transitions, then this can easily be generated by right clicking on the upper state, selecting "Copy with linked items" and then "Paste". (There is more than one way to structure the addition of a state - see the "Different Manifolds" section below for a discussion of this.) Setting the symmetry of the newly created state to Σ+ and manually adjusting the origin of the new state to 27166.8 gives a promising simulation:
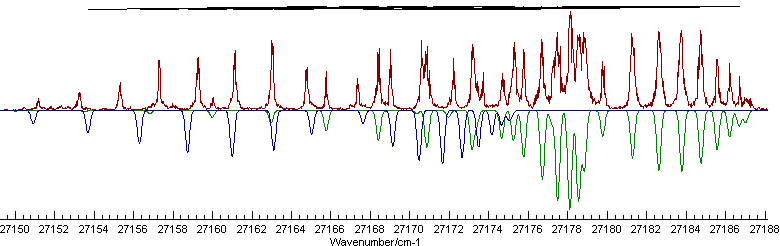
To obtain this appearance the colour of the new state has been
set to "Navy"
and the Π state to "Green";
the file is saved as multc3fitinital.pgo,
and then clicking the "Show
Parts" (![]() ) and "Show Sum" (
) and "Show Sum" (![]() )
buttons. Now, while the simulation is quite good, note that the
indicating marks at the top show a very poor fit, and fitting
with the old line list will fail. Deleting the line list and
starting again would work fine, but there are better ways to do
this which we we will now explore.
)
buttons. Now, while the simulation is quite good, note that the
indicating marks at the top show a very poor fit, and fitting
with the old line list will fail. Deleting the line list and
starting again would work fine, but there are better ways to do
this which we we will now explore.
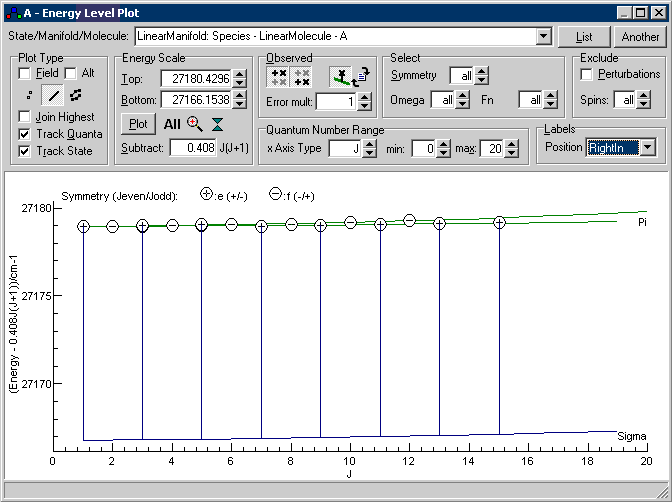
If you have trouble setting this up, the settings are saved
in multc3fitinital.pgo.This
is a plot of energy against J, but with 0.408J(J+1)
subtracted from the energy, so simple states will give
horizontal lines. The two excited states are clearly visible
on this plot as the green and blue horizontal lines, and the
circles give the observed energy levels calculated from the
line list by adding the calculated lower state energy to the
observed transition energy. The vertical lines join the
calculated to the observed energy levels, and it is clear from
this that about half of the assignments are to the wrong
energy level. This is because the line list constructed
specifies states in terms of J, symmetry (+ or -) and
eigenvalue number, and we have just added a state below the
previous one, increasing the eigenvalue number. There are
various ways of dealing with this and we work through several
possibilities here, all of which work for this case, but may
not in more complicated cases.
The first possibility is to put the new state in a different manifold from the initial Π state. The original line list will then work, as the manifold is specified for each state. The disadvantage of this is that perturbations between states can't be simulated, and states close together in energy as these are may well interact. The required set in is shown in the right hand side below, with the original set up on the left:
 |
|
| Setup with two states (Sigma, Pi)
in a single excited state manifold (A) |
Setup with each state in
its own manifold (A for Pi,
Asigma for Sigma) |
Converting between the two is straightforward:
The "Simple" format uses the conventional quantum numbers to
specify the transitions. Such a file can be generated from the
Line List Window using "More,
Save to File, In Simple Format" and choosing a suitable
file name, say APisimple.lin.
Now go to the Log Window, check
than the file you have just created is shown and press "Edit" to edit the
file. A line:
UpperState Pi
needs to be added to the start of the file to identify which
of the two possible upper vibronic states is involved in the
observed transition. The file then needs to be saved - if you
use the Tabslave Editor
included with PGOPHER this step is not necessary as
it a save is forced on each PGOPHER fit or
observation load. The file should look like this:
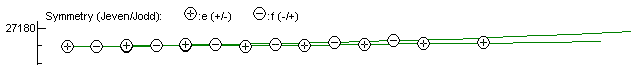
You can check this is right by hitting the reload button in the Energy Level Plot Window and the upper state should now show no long vertical lines, indicating a correct assignment:NQN 2
UpperState Pi
1 e 0 e 27179.774 1 .03172 : LDS751.dr b - C3AX.ovr
3 e 2 e 27181.277 1 .05494 : LDS751.dr b - C3AX.ovr
5 e 4 e 27182.625 1 .0589 : LDS751.dr b - C3AX.ovr
...
 Only two of many possible directives have been used here. Others
include defaultmolecule,
uppermanifold, lowermanifold or lowerstate which might
also be required if there is more than one possibility for each.
For a complete list of directives Line
List Input File Format.
Only two of many possible directives have been used here. Others
include defaultmolecule,
uppermanifold, lowermanifold or lowerstate which might
also be required if there is more than one possibility for each.
For a complete list of directives Line
List Input File Format.An alternative way of specifying transitions is to use
traditional branch notation, such as P(3), implying J'
= 2, J" = 3. Such a file can be generated from the Line List Window using "More,
Save to File, In Branch Format". The resulting file (in
APibranch.lin) looks like
this:
As for the simple format above, the "UpperState Pi" is required to identify which of the two possible upper vibronic states is required. The branch needs to appear without spaces at the beginning of the line - see Branch Format for more details. The rest of the line is as for simple format.UpperState Pi
R(0) 27179.774 1 .03172 : LDS751.dr b - C3AX.ovr
R(2) 27181.277 1 .05494 : LDS751.dr b - C3AX.ovr
R(4) 27182.625 1 .0589 : LDS751.dr b - C3AX.ovr
...
The final option in fixing the line list is simply to change
the eigenvalue numbers that are wrong in the new simulation
set up. In the line list window this will be the #' column,
and in this case the values for the odd J levels need
increasing by 1. This change could be done manually, but a
further change may need making if a state is added or removed,
so we use the indexoffsets
directive. To use this we put the line list in a separate
file. Select "More, Save to File, In Standard Format"
in the Line List Window window and
choose a suitable filename, say APi.lin. To add the
required offsets add:
upperindexoffsets 1 0
at the start of the file and then save the changed file. You
can check this is right by hitting the reload button in the Energy Level Plot Window and the
upper state should now show no long vertical lines, indicating
a correct assignment:

As an alternative the quantum numbers given in the comments
at the end of each line can also be used to work out the
eigenvalue number . In the Line List
Window this can be forced for an individual line by
clearing the eigenvalue number (#' and #") columns or in a
line list file by setting the eigenvalue number to 0. The
symmetry (S', S") can also be similarly left unset, though a
negative value, rather than 0 must be used in the line list
file to flag this. To force this for the entire file use:
upperindexoffsets search
to work out the eigenvalue numbers or
upperindexoffsets searchall
to work out both symmetry and eigenvalue number. Alternatively:
UpperState Pi
can be used to indicate that the eigenvalue numbers are to be
taken within the state. Any of these methods will work fine in
many cases, though where the quantum numbers are ambiguous,
typically in the presence of strong state mixing, the quantum
numbers assigned can change as the parameters change. In
fitting the quantum number assignments for any given state can
flip between successive fit cycles.
A few lines of the line list file are shown below; see Line List Input File Format for a
detailed explanation of each field.
LinearMolecule A 1 1 1 X 0 0 1 27179.774 1 .03172 0 : R(0) : A Pi 1 e - X v=0 0 e : LDS751.dr b - C3AX.ovr
LinearMolecule A 3 1 1 X 2 0 1 27181.277 1 .05494 0 : R(2) : A Pi 3 e - X v=0 2 e : LDS751.dr b - C3AX.ovr
LinearMolecule A 5 1 1 X 4 0 1 27182.625 1 .0589 0 : R(4) : A Pi 5 e - X v=0 4 e : LDS751.dr b - C3AX.ovr
...
Notes on this file:
UpperManifold A
1 1 1 0 0 1 27179.774 1
3 1 1 2 0 1 27181.277 1
5 1 1 4 0 1 27182.625 1
...
NQN 2
1 e 2 e 27164.765 1
3 e 4 e 27163.026 1
...
To use this file with one of the previous Π state files
set up a separate file, say Aboth.lin,
with the following format:
Colour Green
Include APisimple.lin
Colour Navy
UpperState Sigma
Include ASigma.lin
The include statements simply read the contents of the given file. All the information could be put in one file, or split in any other way as convenient. The colour directives set the colour of the points in the Residuals Window and the indicating marks at the top of the main window. The final fit gives an excellent overall simulation: